Abstract
Due to potentially hostile behaviors and elusive habitats, moray eels (Muraenidae) as one group of apex predators in coral reefs all across the globe have not been well investigated. Here, we constructed a chromosome-level genome assembly for the representative Reeve’s moray eel (Gymnothorax reevesii). This haplotype genome assembly is 2.17 Gb in length, and 97.87% of the sequences are anchored into 21 chromosomes. It contains 56.34% repetitive sequences and 23,812 protein-coding genes, of which 96.77% are functionally annotated. This sequenced marine species in Anguilliformes makes a good complement to the genetic resource of eel genomes. It not only provides a genetic resource for in-depth studies of the Reeve’s moray eel, but also enables deep-going genomic comparisons among various eels.
Similar content being viewed by others
Background & Summary
Fish in family Muraenidae, commonly referred to as moray eels, are one group of the most species-rich of the order Anguilliformes. At present, approximately 210 species are recognized1,2. They are widely distributed in tropical, subtropical, and temperate waters worldwide, particularly in the region of Indo-Pacific3,4,5. Muraenidae species are cryptic, primarily found in shallow waters as rock and coral reef inhabitants, which play crucial roles in the maintenance of coral reef biodiversity1. They exhibit a wide range of color patterns, from uniform distinctive patterns of spots to blotches, bars, and reticulations. In a number of coastal countries, Muraenidae species are of commercial importance to the fishery industry due to their high nutritional value. Besides, their spectacular external coloration makes them well-liked in marine aquariums. To date, moray eels are not well-researched due to their cryptic habitats and irregularly aggressive behaviors6.
Anguilliformes species are scale-less with elongated bodies and usually lack pelvic fins7,8,9. Unlike other families in this order, some Muraenidae species don’t even have pectoral fins, and their opercular bones are degenerated with only small and round branchial pores opening outside2. Moray eels also have an unexpectedly wide variety of body sizes as per the whole length and mass. Their adults range in body mass from 4 to above 600 g, and in total length from 10 cm to 400 cm. Remarkably, the average number of vertebral numbers in morays varies by 2.4 folds, which is equal to the variance among extant snakes10. The maximum length and body shape are two parameters for diversifying among organisms, as each holds an effect on organismal-environment interactions11,12. Due to their distinctive morphological features, moray eels can be utilized as a good model for thoroughly exploring molecular mechanisms of evolutionary body patterns.
Both freshwater and marine eels differ a lot in phenotypic evolution, adaptation, and speciation and are closely related to chromosome evolution including rearrangements as has been proved by previous studies13. The majority of freshwater eels have only 19 pairs of chromosomes14, but some marine eels such as leopard moray eel (Enchelycore pardalis) and brown moray eel (Gymnothorax unicolor) have 21 pairs15. An in-depth investigation of this chromosomal difference between these eels may be instructive to reveal phenotypic, adaptive, and speciation differences between the two groups.
Previous studies have primarily concentrated on the morphology, taxonomy, and germplasm resource of moray eels, while limited genetic and genomic resources have largely constrained the conservation as well as utilization of these commercial species. Reeve’s moray eel (Gymnothorax reevesii), as a representative species in the family Muraenidae, has an elongated body and does not have pectoral and pelvic fins (Fig. 1). G. reevesii is a subtropical and tropical species in the Northwest Pacific, and most distribute from southern Japan to the South China Sea16. Its body color of yellowish brown to brownish, with numerous large obscure dark brownish spots in 3–5 rows along the body side. Among Muraenidae fishes, G. reevesii is a common catch in coastal areas of eastern and southern China with commercial importance to local fisheries17. In recent years it is also tamed as cultured fish and becomes one of the few Muraenidae species that can be artificially cultured with a vast developmental and economic potential. But until now very few studies have been focused on this species18, indicating that a huge research gap is still existed, and thereby further information about this species is needed to be explored. In our present study, a chromosome-level genome assembly of G. reevesii was constructed using an integrated strategy of MGI19 (second generation), PacBio (third generation), and Hi-C sequencing technologies, and we then characterized the high repeat content in the genome, inferred the phylogeny, and performed a chromosomal synteny analysis. This sequenced marine species in Anguilliformes makes a good complement to the genetic resource of eel genomes.
Images of the sequenced Reeve’s moray eel. (a) A lateral full-body view of the fish. (b) A thoracic side view of the fish. The arrow points to the gill apertures. The pelvic and pectoral fins are not visible in the field of this view. (c) A caudal ventral view of the fish. The dashed circle covers the cloacal area. The arrow points to where the caudal fin begins. (d) A lateral view of the fish’s tail. The caudal fin is not seen in the area pointed by the arrow. (e–g) An enlarged view of the thorax, ventral, and tail of the fish. Obviously, no scales are visible on the skin surface of these areas.
Methods
Sample collection
A female Reeve’s moray eel (Fig. 1a) was collected from Daya Bay Aquatic Testing Center in Huiyan District, Huizhou City, Guangdong Province, China. We pooled the muscle (about 5 g) of this individual for whole genome sequencing (short-read, long-read, and Hi-C sequencing), and the muscle, gonads, brain, and liver (50 mg for each tissue) for transcriptome sequencing. These samples were cut into small pieces and freshly frozen in liquid nitrogen, and then were stored at −80 °C until use. The Animal Ethics Committee of Zhongkai University of Agriculture and Engineering (Guangzhou, China) approved our sampling pipeline.
DNA extraction and genome sequencing
Extraction and purification of genomic DNA (gDNA) from the muscle (2 g) was carried out using a blood & cell culture DNA kit (Qiagen, USA) in accordance with the manufacturer’s protocol.
The extracted gDNA (1.5 ug subsample) was randomly fragmented and used to create a 350-bp insert-size library by using MGIEasy universal DNA library prep set (MGI, China) for subsequent sequencing on a MGISEQ 2000 platform (MGI, China). A total of 65.39 Gb of paired-end raw reads (150 bp in length) were generated and then filtered via the SOAPfilter v2.220 (default parameters) to remove adaptor sequences and low-quality reads. We finally obtained approximately 60.68 Gb of clean reads for estimation of the genome size and subsequent assembling.
Furthermore, we sampled 2 ug gDNA to construct long-read libraries by using a SMRTbell Express Template Prep Kit 2.0 for HiFi sequencing based on PacBio’s standard protocol (Pacific Biosciences, USA), which were sequenced through a PacBio Sequel II System. CCS software (SMRT Link v9.0)21 was then applied to generate the consensus sequences (-min-passes 1 --min-rq 0.99 --min-length 100). About 5.46 million consensus reads (68.88 Gb) with a mean length of 12.62 kb were obtained.
For the Hi-C sequencing, muscle tissue (about 1 g) from the same individual was collected, and DNA libraries were constructed by using GrandOmics Hi-C kit (the applied restriction enzyme is DpnII; GrandOmics, China) according to the manufacturer’s protocol. The Hi-C libraries were then sequenced on an Illumina Novaseq platform (Illumina, USA). In total, 305.18 Gb of Hi-C paired-end raw reads (150 bp in length) were generated. Subsequently, fastp22 was applied to filter the adaptors, and those reads shorter than 30 bp or of low-quality (quality scores < 20). Finally, 99.98% reads (304.92 Gb) were retained for construction of pseudo-chromosomes.
RNA extraction and transcriptome sequencing
RNA samples were extracted from muscle, gonad, brain, and liver tissues (50 mg for each tissue) using a standard Trizol protocol (Invitrogen, USA), and purified using a Qiagen RNeasy mini kit (Qiagen, USA). RNA with equal amounts from each tissue was mixed for creating an Illumina cDNA library followed the manufacture’s guideline, which was then sequenced on a HiSeq X Ten platform (Illumina, USA). Around 6.01-Gb transcriptome data were generated for assistance to genome and gene annotations.
Genome-size estimation
To estimate the genome size of Reeve’s moray eel, a k-mer analysis23 was performed by using MGI clean reads. Through the k-mer counting (KMC) program24 and genome character estimator (GCE) software25, the 17-mer frequency was calculated. The genome size was then estimated by assessing the 17-mer depth distribution through the equation of G = K_num/K_depth (G is the genome size, K_depth represents the k-mer depth, and K_num stands for the total number of 17-mers). The estimated genome size of Reeve’s moray eel is therefore about 2.05 Gb, and the genomic heterozygosity rate was predicted to be 1.02% (Fig. 2a).
A chromosome-level genome assembly of the Reeve’s moray eel. (a) A GenomeScope k-mer plot. (b) A total of 21 distinct blocks were visualized in the Hi-C contact matrixes. (c) A Circos plot summarizing the genome features. From outside to inside: (I) the 21 pseudo-chromosomes, (II) gene density, (III) repeats, and (IV) GC content. Links inside the Circos refer to internal syntenic blocks among various chromosomes within the assembled genome.
De novo genome assembly
After obtaining subreads, the PacBio long reads were de novo assembled into contigs through hifiasm (v0.16.0)26 with default parameters. These contigs were then polished with Nextpolish (v1.10)27 using the MGI short reads to fix possible base errors. The primary genome assembly was 2.23 Gb in length, consistent with the estimated genome size.
We employed multiple methods to evaluate the quality of this assembly. First, Merqury v1.328 shows that the estimated completeness, QV, and error rate of the assembly were 86.33%, 41.47 and 7.12e-05, respectively. A k-mer spectra plot is provided (Fig. 3a). Second, Benchmarking Universal Single Copy Orthologs (BUSCO) v5.2.229 against actinopterygii_odb10 database was employed to assess the completeness of this genome assembly, showing that the assembly contains 94.89% of complete BUSCO genes including 88.3% single-copies and 6.59% duplicates (Table 1), suggesting that this genome assembly is of high quality. Third, Core Eukaryotic Gene Mapping Approach (CEGMA v2.5)30 also shows that 223 (89.92%) out of 248 core genes were successfully assembled (Fig. 3b). Finally, we mapped the long reads back to the assembly via minimap2 v2.1531, and calculated the average GC content and sequencing depth over 10-kb windows, which shows that the assembled genome is clean without contamination (Fig. 3c). Gfastats v1.3.132 was applied to obtain assembly summary statistics (see more details in Table 1).
Quality assessment of the genome assembly of Reeve’s moray eel. (a) Merqury spectrum plot for the genome assembly. K-mers in the assembly (red) and only in the reads (gray) are presented. (b) Genome completeness assessment by CEGMA presenting the annotated core genes in the assembly. Green bars denote the complete genes, while light green bars denote the complete and partial genes combined. (c) GC-depth plot. The x- and y-axis represent GC content and sequencing depth respectively, and corresponding histograms are on the top and right.
Pseudo-chromosome construction
Based on this high-quality genome assembly, Hi-C technique was subsequently employed to construct pseudo-chromosomes for the Reeve’s moray eel. First, Hi-C clean reads were mapped to the assembled contigs using bowtie2 (v2.3.2)33 (-end-to-end --very-sensitive -L 30). Subsequently, HiC-Pro (v2.8.1)34 pipeline was applied to detect valid ligation products and only valid contact paired reads were retained for further analysis. Based on these valid reads, the primary assembly was oriented, ordered, and clustered onto chromosomes through LACHESIS35, with optimized parameters (CLUSTER_MIN_RE_SITES = 100, CLUSTER NONINFORMATIVE RATIO = 1.4, CLUSTER_MAX_LINK_DENSITY = 2.5, ORDER MIN N RES IN SHREDS = 60, ORDER MIN N RES IN TRUNK = 60). JuiceBox v1.11.0836 was employed for manual correction of placement and orientation mistakes with glaring distinct chromatin interaction patterns. We hence obtained the final genome assembly with a size of 2.17 Gb, of which 97.87% are anchored into 21 chromosomes (Fig. 2). The scaffold and contig N50 values of the overall chromosome-level genome assembly are 112.89 Mb and 53.38 Mb, respectively, reaching a relatively high level among sequenced fish species.
High repeat content in the Reeve’s moray eel genome
Transposable elements (TE) in the G. reevesii genome were predicted by combination of homology-based and ab initio predictions. In general, default settings from RepeatModeler37 and MITE-Hunter38 were employed to obtain an ab initio repetition library, which was then aligned to Repbase39 with TEclass tool40 for classifying the details of each repeat family. By mapping sequences against the Repbase TE library and de novo repeat library, RepeatMasker37 was applied to check known and new TEs so as to further identify these repeats across the assembled genome. Finally, a total of 1.23 Gb of repetitive sequences were annotated in the assembly, including 1.09 Gb of TEs (Table 2).
In summary, repetitive sequences account for up to 56.34% of the genome of Reeve’s moray eel, including 50.15% of TEs (Table 2), which content is much higher than those of freshwater eels14 and most sequenced fishes so far that usually have a TE content less than 45%41. We then detected the repeat content in each chromosome of the Reeve’s moray eel and the freshwater eels. A plot of the repeat distribution throughout corresponding chromosomes, drawn by the RIdeogram package42, indicated that each chromosome of the Reeve’s moray eel also has a higher repeat content when compared to those freshwater counterparts (Fig. 4a).
The Reeve’s moray eel had a higher content of repetitive sequences than freshwater eels. (a) Repeat density throughout the chromosomes of the examined eels. The redder indicates the higher repeat content, while the bluer marks the lower repeat content. (b) Distribution of the four major TE types. Red, black, yellow, and green bars represent DNA, LINE, LTR, and SINE repeats, respectively.
More specifically, the Reeve’s moray eel genome has 50.15% of TEs, accounting for 89.01% of the total repetitive sequences. The combined length of the sequences representing each type of DNA, LTR, LINE, and SINE are 560.92, 200.04, 165.14, and 20.60 kb, respectively, which shows remarkable expansions of DNA and LTR sequences in the genome of Reeve’s moray eel compared to the freshwater eels (Fig. 4b). Considering that all these eel genomes with Mb-level contig N50 values were assembled from long reads with sufficient coverage, this difference appears be real among genomes and not the result of unassembled repeats in the freshwater eel genomes.
Gene annotation and functional assignment
In the repeat-masked genome, gene prediction was carried out using three methods, including homology, transcriptome-based, and ab initio annotations. For the homology prediction, GeMoMa43 was employed to align homologous proteins from relevant species (including Japanese eel Anguilla japonica, spotted gar Lepisosteus oculatus, zebrafish Danio rerio, three-spined stickleback Gasterosteus aculeatus, and Fugu Takifugu rubripes) to our assembly to predict gene structures. For the transcriptome-based gene prediction, clean transcriptome reads were mapped to the assembled genome by STAR44. PASA45 was then applied to predict open reading frames (ORFs) and stringtie46 was employed to assemble transcripts for gene structure annotation. For the ab initio annotation, transcriptome reads were de novo assembled by Trinity (v2.13.2)47, and 300 full-length transcripts were randomly selected to build a library as a training set. Based on this training set, Augustus48 with default parameters was then employed to predict genes. Finally, EVidenceModeler (EVM)45 was applied to create a combined and non-redundant gene set after removal of those miscoded genes, and the final protein sequences were deduced by standard genetic codes.
Motifs/domains and gene functions of these annotated genes were predicted by blasting (BLASTp) the deduced protein sequences against various public databases, including SwissProt49, NCBI NR, KEGG50, KOG51, and Gene Ontology52 (GO), with an E-value cutoff of 1e−05.
Finally, in the assembled genome of Reeve’s moray eel, a total of 23,812 protein-coding genes were annotated, with an average gene length of 36,705.61 bp, an average coding sequence (CDS) length of 1,738.42 bp, and an average number of 8.51 exons per gene. Among these genes, 96.77% (23,812 genes) were annotated in the searched protein databases (Table 3). In addition, the CDSs were further translated into protein sequences with standard genetic codes, followed by self-alignment using BLASTp for construction of internal syntenic blocks by MCscan53 with parameters “-a -e 1e-5 -u 1 -s 8”. A Circos54 plot (Fig. 2c) was then generated to show (I) lengths of 21 pseudo-chromosomes, (II) gene density (percentage of genes per 100-kb window), (III) repeat density (minimum 0%, maximum 100%), (IV) GC content (minimum 30%, maximum 65%), and internal syntenic blocks (in the center).
Phylogenetic tree
A species tree was constructed using those single-copy orthologs from the whole genomes of Reeve’s moray eel and other 12 representative ray-finned species, with ropefish (Erpetoichthys calabaricus) as the outgroup. Both the Maximum Likelihood tree (constructed using PhyML55) and the Bayesian tree (inferred by MrBayes56) present the same topology with 100% for the node bootstrap values or 1 for the node support values. The consensus tree was further time-calibrated using MCMCTREE program in the PAML package57. The tree shows that Reeve’s moray eel and two examined freshwater eels make up the Anguilliformes clade, forming a sister group to the Elopiformes composed of two tarpons, and that Reeve’s moray eel diverged from both freshwater eels 101.84 million years ago (Mya; Fig. 5). These data are consistent with previous reports58,59,60.
A phylogenetic tree of G. reevesii and other representative ray-finned fish species. Number at each node means the estimated divergence time. Blue dots denote the fossils used for time calibration. The Anguilliformes lineage is highlighted in a yellow shadow. The pair numbers of chromosomes are 21 and 19 for the Reeve’s moray eel and freshwater eels, respectively.
Data Records
The genome assembly and raw reads of the genome and transcriptome sequencing for Reeve’s moray eel were deposited at NCBI under the accession number PRJNA93405561. Raw reads are available in the Sequence Reads Archive (SRA) with the accession number SRP430070. The genome assembly was deposited at GenBank with the accession number GCA_029721435.162. Repeat and gene annotation files are publicly available in FigShare depository with accessions https://doi.org/10.6084/m9.figshare.2235798763 and https://doi.org/10.6084/m9.figshare.2235820964, respectively.
Technical Validation
Quality of extracted gDNA was detected by agarose gel electrophoresis (a main band at 20 kb) and Nanodrop spectrophotometer (Thermo Fisher Scientific, USA). The quality of the isolation met the standards that OD260/280 value ranges from 1.8 to 2.0, and OD260/230 is between 2.0–2.2. Quality of extracted RNA was examined by a 2100 Bioanalyzer (RIN > 7.0, 28 S/18 S > 1.0; Agilent Technologies, USA).
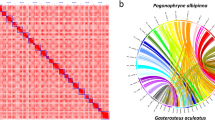
Reeve’s moray eel has 21 pairs of chromosomes (Chr) whereas most freshwater eels have only 19 pairs, which is evidenced by both karyotypic observation and genomic sequencing14,65,66,67. We therefore performed a chromosomal synteny analysis using the JCVI package68. Our plot shows that European eel and American eel had perfect one-to-one correspondences among their chromosomes (Fig. 5), and Reeve’s moray eel showed a generally conserved chromosomal synteny although with a few large fragmental exchanges. For example, both Chr2 and Chr3 in American eel and European eel correspond to two chromosomes (10 & 20, and 14 & 18, respectively) of Reeve’s moray eel. In addition, part of the Chr5 fused to the end of the Chr13 (American eel) or Chr12 (European eel) of the freshwater eels, forming Chr2 in the Reeve’s moray eel, and the other half of the Chr5 was preserved as a single chromosome (Chr17) in moray eel, presenting a two-to-two synteny pattern (Fig. 6). The chromosome synteny analysis implies that there is generally conserved chromosomal synteny between moray eel and freshwater eels, but a few fissions and fusions happened after their divergence, leading to 21 and 19 pairs of chromosomes in the two lineages respectively14.
Code availability
The versions and parameters of bioinformatic tools used in this study have been described in the Method section. If no parameter is provided, the default is used. No custom code was used.
References
Böhlke, E. B. & Smith, D. G. Type catalogue of Indo-Pacific Muraenidae. Proc. Acad. Nat. Sci. Phila. 152, 89–172 (2002).
Smith, D. G., Bogorodsky, S. V., Mal, A. O. & Alpermann, T. J. Review of the moray eels (Anguilliformes: Muraenidae) of the Red Sea, with description of a new species. Zootaxa 4704, 1–87 (2019).
Smith, D. G. A checklist of the moray eels of the world (Teleostei: Anguilliformes: Muraenidae). Zootaxa 3474, 1–64 (2012).
Nelson, J. S., Grande, T. C. & Wilson, M. V. H. Fishes of the World. (John Wiley & Sons, 2016).
Mohapatra, A., Mohanty, S. R., Smith, D. G., Mishra, S. S. & Roy, S. Gymnothorax odishi sp. nov. (Muraenidae: Muraeninae), a short brown unpatterned moray eel from Bay of Bengal, India. Zootaxa 4420, 123–130 (2018).
Reece, J. S., Bowen, B. W., Smith, D. G. & Larson, A. Molecular phylogenetics of moray eels (Muraenidae) demonstrates multiple origins of a shell-crushing jaw (Gymnomuraena, Echidna) and multiple colonizations of the Atlantic Ocean. Mol. Phylogenet. Evol. 57, 829–835 (2010).
Ramos-Castro, M., Loh, K. H. & Chen, H. M. A descriptive and comparative neurocranium morphology of Anguilliformes fishes in Taiwan waters. Zootaxa 5023, 509–536 (2021).
Arai, T. Ecology and evolution of migration in the freshwater eels of the genus Anguilla Schrank, 1798. Heliyon 6, e05176 (2020).
López, J. A., Westneat, M. W. & Hanel, R. The phylogenetic affinities of the mysterious Anguilliform genera Coloconger and Thalassenchelys as supported by mtDNA sequences. Copeia 2007, 959–966 (2007).
Reece, J. S. & Mehta, R. S. Evolutionary history of elongation and maximum body length in moray eels (Anguilliformes: Muraenidae). Biol. J. Linn. Soc. 109, 861–875 (2013).
Brown, J. H. & Lasiewski, R. C. Metabolism of weasels: the cost of being long and thin. Ecology 53, 939–943 (1972).
Schmidt-Nielsen, K. & Knut, S.N. Scaling: why is animal size so important? (Cambridge university press, 1984).
Damas, J., Corbo, M. & Lewin, H. A. Vertebrate chromosome evolution. Annu. Rev. Anim. Biosci. 9, 1–27 (2021).
Wang, H. et al. A Chromosome-level assembly of the Japanese eel genome, insights into gene duplication and chromosomal reorganization. Gigascience 11, giac120 (2022).
Ronchetti, E., Salvadori, S. & Deiana, A. M. Genome size and AT content in Anguilliformes. Eur. J. Histochem. 39, 259–264 (1995).
Prokofiev, A. M. Additions to the species composition of morays of the Bay of Nha Trang (South China Sea, central Vietnam)(Anguilliformes: Muraenidae). J. Ichthyol. 50, 38–43 (2010).
Chen, H. M., Shao, K. T. & Chen, C. T. A review of the muraenid eels (Family Muraenidae) from Taiwan with descriptions of twelve new records. Zool. Stud. 33, 44–64 (1994).
Ramos-Castro, M., Chen, H. M. & Mao, B. S. Descriptions of neurocranium morphology in 34 species of moray eels (Muraenidae) found in Taiwan. Zootaxa 4810, 081–109 (2020).
Blake, J. A., Bult, C. J., Kadin, J. A., Richardson, J. E. & Eppig, J. T. The Mouse Genome Database (MGD): premier model organism resource for mammalian genomics and genetics. Nucleic Acids Res. 39, D842–D848 (2011).
Luo, R. et al. SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. Gigascience 1, 18 (2012).
Rhoads, A. & Au, K. F. PacBio Sequencing and Its Applications. Genom. Proteom. Bioinform. 13, 278–289 (2015).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Vurture, G. W. et al. GenomeScope: fast reference-free genome profiling from short reads. Bioinformatics 33, 2202–2204 (2017).
Deorowicz, S., Kokot, M., Grabowski, S. & Debudaj-Grabysz, A. KMC 2: fast and resource-frugal k-mer counting. Bioinformatics 31, 1569–1576 (2015).
Sun, H., Ding, J., Piednoël, M. & Schneeberger, K. findGSE: estimating genome size variation within human and Arabidopsis using k-mer frequencies. Bioinformatics 34, 550–557 (2018).
Cheng, H., Concepcion, G. T., Feng, X., Zhang, H. & Li, H. Haplotype-resolved de novo assembly using phased assembly graphs with hifiasm. Nat. Methods 18, 170–175 (2021).
Hu, J., Fan, J., Sun, Z. & Liu, S. NextPolish: a fast and efficient genome polishing tool for long-read assembly. Bioinformatics 36, 2253–2255 (2020).
Rhie, A., Walenz, B. P., Koren, S. & Phillippy, A. M. Merqury: reference-free quality, completeness, and phasing assessment for genome assemblies. Genome Biol. 21, 245 (2020).
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212 (2015).
Parra, G., Bradnam, K. & Korf, I. CEGMA: a pipeline to accurately annotate core genes in eukaryotic genomes. Bioinformatics 23, 1061–1067 (2007).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Formenti, G. et al. Gfastats: conversion, evaluation and manipulation of genome sequences using assembly graphs. Bioinformatics 38, 4214–4216 (2022).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Servant, N. et al. HiC-Pro: an optimized and flexible pipeline for Hi-C data processing. Genome Biol. 16, 259 (2015).
Burton, J. N. et al. Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions. Nat. Biotechnol. 31, 1119–1125 (2013).
Durand, N. C. et al. Juicebox provides a visualization system for Hi-C contact maps with unlimited zoom. Cell Syst. 3, 99–101 (2016).
Chen, N. Using RepeatMasker to identify repetitive elements in genomic sequences. Curr. Protoc. Bioinformatics 5, 4.10.1–4.10.14 (2004).
Han, Y. & Wessler, S. R. MITE-Hunter: a program for discovering miniature inverted-repeat transposable elements from genomic sequences. Nucleic Acids Res. 38, e199 (2010).
Jurka, J. et al. Repbase Update, a database of eukaryotic repetitive elements. Cytogenet. Genome Res. 110, 462–467 (2005).
Abrusán, G., Grundmann, N., DeMester, L. & Makalowski, W. TEclass—a tool for automated classification of unknown eukaryotic transposable elements. Bioinformatics 25, 1329–1330 (2009).
Shao, F., Han, M. & Peng, Z. Evolution and diversity of transposable elements in fish genomes. Sci. Rep. 9, 15399 (2019).
Hao, Z. et al. RIdeogram: drawing SVG graphics to visualize and map genome-wide data on the idiograms. PeerJ Comput. Sci. 6, e251 (2020).
Keilwagen, J. et al. Using intron position conservation for homology-based gene prediction. Nucleic Acids Res. 44, e89 (2016).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Haas, B. J. et al. Automated eukaryotic gene structure annotation using EVidenceModeler and the program to assemble spliced alignments. Genome Biol. 9, R7 (2008).
Kovaka, S. et al. Transcriptome assembly from long-read RNA-seq alignments with StringTie2. Genome Biol. 20, 278 (2019).
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 29, 644–652 (2011).
Stanke, M., Diekhans, M., Baertsch, R. & Haussler, D. Using native and syntenically mapped cDNA alignments to improve de novo gene finding. Bioinformatics 24, 637–644 (2008).
Bairoch, A. et al. The universal protein resource (UniProt). Nucleic Acids Res. 33, D154–D159 (2005).
Ogata, H. et al. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 27, 29–34 (1999).
Galperin, M. Y., Makarova, K. S., Wolf, Y. I. & Koonin, E. V. Expanded microbial genome coverage and improved protein family annotation in the COG database. Nucleic Acids Res. 43, D261–D269 (2015).
The Gene Ontology Consortium. The Gene Ontology resource: 20 years and still GOing strong. Nucleic Acids Res. 47, D330–D338 (2019).
Wang, Y. et al. MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 40, e49 (2012).
Krzywinski, M. et al. Circos: an information aesthetic for comparative genomics. Genome Res. 19, 1639–1645 (2009).
Guindon, S. et al. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst. Biol. 59, 307–321 (2010).
Ronquist, F. et al. MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 61, 539–542 (2012).
Yang, Z. H. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007).
Tang, K. L. & Fielitz, C. Phylogeny of moray eels (Anguilliformes: Muraenidae), with a revised classification of true eels (Teleostei: Elopomorpha: Anguilliformes). Mitochondrial DNA 24, 55–66 (2013).
Barth, J. M. I. et al. Stable species boundaries despite ten million years of hybridization in tropical eels. Nat. Commun. 11, 1433 (2020).
Rabosky, D. L. et al. Rates of speciation and morphological evolution are correlated across the largest vertebrate radiation. Nat. Commun. 4, 1958 (2013).
NCBI Bioproject https://www.ncbi.nlm.nih.gov/bioproject/PRJNA934055 (2023).
NCBI Genbank https://identifiers.org/ncbi/insdc.gca:GCA_029721435.1 (2023).
Huang, L. Repeats annotation. figshare https://doi.org/10.6084/m9.figshare.22357987.v1 (2023).
Huang, L. Gymnothorax reevesii gene annotation (gff, cds, pep). figshare https://doi.org/10.6084/m9.figshare.22358209.v1 (2023).
Park, E. H. & Kang, Y. S. Karyotype conservation and difference in DNA amount in Anguilloid fishes. Science 193, 64–66 (1976).
Vasconcelos, A. J. M. & Molina, W. F. Cytogenetical studies in five Atlantic Anguilliformes fishes. Genet. Mol. Biol. 32, 83–90 (2009).
Kai, W. et al. A ddRAD-based genetic map and its integration with the genome assembly of Japanese eel (Anguilla japonica) provides insights into genome evolution after the teleost-specific genome duplication. BMC Genomics 15, 233 (2014).
Tang, H. et al. Synteny and collinearity in plant genomes. Science 320, 486–488 (2008).
Acknowledgements
This project was supported by Guangdong Modern Agricultural Industry Technology Innovation Team Special Project (no. 2019KJ140), National Key Research and Development Program of China (no. 2022YFE0139700), and Science and Technology Plan Project of Qingyuan City (no. 2022KJJH064).
Author information
Authors and Affiliations
Contributions
L.L. and Q.S. conceived this project. K.Z., Y.H., Y.X.Z. and R.S.L. participated in data analysis. Y.X.Z., R.S.L., J.H.W., Q.Q.L. and Y.N.C. collected the samples. K.Z., Y.H. and C.B. assembled the genome. R.H.L. and X.M.Z. generated the annotation set. K.Z., Y.H., R.S.L., Y.X.Z. and L.L. wrote the manuscript. Q.S. and L.L. revised the manuscript. All authors have read and approved the final manuscript for publication.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, K., Huang, Y., Zhang, Y. et al. A chromosome-level reference genome assembly of the Reeve’s moray eel (Gymnothorax reevesii). Sci Data 10, 501 (2023). https://doi.org/10.1038/s41597-023-02394-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-023-02394-7
This article is cited by
-
Chromosome-level genome assembly and annotation of eel goby (Odontamblyopus rebecca)
Scientific Data (2024)